What protein (enzyme) is used to produce ATP?
| ATP Synthase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
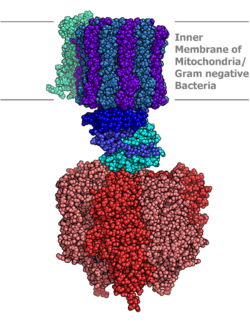 Molecular model of ATP synthase determined by 10-ray crystallography. Stator is not shown hither. | |||||||||
| Identifiers | |||||||||
| EC no. | 7.1.2.2 | ||||||||
| CAS no. | 9000-83-three | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | contour | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
ATP synthase is a protein that catalyzes the formation of the free energy storage molecule adenosine triphosphate (ATP) using adenosine diphosphate (ADP) and inorganic phosphate (Pi). It is classified nether ligases as information technology changes ADP by the germination of P-O bail (phosphodiester bond). ATP synthase is a molecular machine. The overall reaction catalyzed by ATP synthase is:
- ADP + Pi + 2H+ out ⇌ ATP + HiiO + 2H+ in
The formation of ATP from ADP and Pi is energetically unfavorable and would commonly keep in the contrary management. In order to drive this reaction forward, ATP synthase couples ATP synthesis during cellular respiration to an electrochemical gradient created by the difference in proton (H+) concentration across the inner mitochondrial membrane in eukaryotes or the plasma membrane in bacteria. During photosynthesis in plants, ATP is synthesized by ATP synthase using a proton gradient created in the thylakoid lumen through the thylakoid membrane and into the chloroplast stroma.
Eukaryotic ATP synthases are F-ATPases, running "in contrary" for an ATPase. This article deals mainly with this type. An F-ATPase consists of 2 main subunits, FO and F1, which has a rotational motor mechanism allowing for ATP production.[1] [2]
Classification [edit]
The Fone fraction derives its proper noun from the term "Fraction 1" and FO (written as a subscript letter "o", not "goose egg") derives its name from being the binding fraction for oligomycin, a blazon of naturally derived antibiotic that is able to inhibit the FO unit of measurement of ATP synthase.[3] [iv] These functional regions consist of unlike poly peptide subunits — refer to tables. This enzyme is used in synthesis of ATP through aerobic respiration.
Structure and office [edit]

Bovine mitochondrial ATP synthase. The FO, F1, axle, and stator regions are colour coded magenta, green, orange, and cyan respectively.[5] [half-dozen]
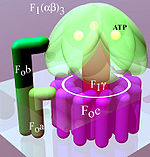

Simplified model of FOF1-ATPase alias ATP synthase of Eastward. coli. Subunits of the enzyme are labeled accordingly.
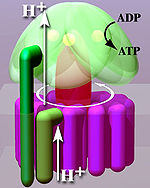
Rotation engine of ATP synthase.
Located within the thylakoid membrane and the inner mitochondrial membrane, ATP synthase consists of two regions FO and F1. FO causes rotation of F1 and is made of c-ring and subunits a, two b, F6. F1 is made of α, β, γ, and δ subunits. Fi has a water-soluble part that can hydrolyze ATP. FO on the other paw has mainly hydrophobic regions. FO Fone creates a pathway for protons motility beyond the membrane.[7]
Fone region [edit]
The F1 portion of ATP synthase is hydrophilic and responsible for hydrolyzing ATP. The F1 unit protrudes into the mitochondrial matrix space. Subunits α and β make a hexamer with 6 binding sites. Three of them are catalytically inactive and they bind ADP.
Three other subunits catalyze the ATP synthesis. The other Fi subunits γ, δ, and ε are a part of a rotational motor machinery (rotor/axle). The γ subunit allows β to get through conformational changes (i.e., closed, half open up, and open states) that allow for ATP to be bound and released once synthesized. The Fi particle is large and can be seen in the transmission electron microscope by negative staining.[8] These are particles of 9 nm diameter that pepper the inner mitochondrial membrane.
| Subunit | Man Factor | Notation |
|---|---|---|
| alpha | ATP5A1, ATPAF2 | |
| beta | ATP5B, ATPAF1 | |
| gamma | ATP5C1 | |
| delta | ATP5D | Mitochondrial "delta" is bacterial/chloroplastic epsilon. |
| epsilon | ATP5E | Unique to mitochondria. |
| OSCP | ATP5O | Called "delta" in bacterial and chloroplastic versions. |
FO region [edit]

FO subunit F6 from the peripheral stalk region of ATP synthase.[10]
FO is a water insoluble protein with eight subunits and a transmembrane ring. The ring has a tetramer shape with a helix loop helix protein that goes through conformational changes when protonated and deprotonated, pushing neighboring subunits to rotate, causing the spinning of FO which and then also affects conformation of F1, resulting in switching of states of alpha and beta subunits. The FO region of ATP synthase is a proton pore that is embedded in the mitochondrial membrane. It consists of three primary subunits, a, b, and c. Six c subunits make up the rotor band, and subunit b makes up a stalk connecting to Fane OSCP that prevents the αβ hexamer from rotating. Subunit a connects b to the c ring.[11] Humans have 6 additional subunits, d, e, f, g, F6, and viii (or A6L). This part of the enzyme is located in the mitochondrial inner membrane and couples proton translocation to the rotation that causes ATP synthesis in the F1 region.
In eukaryotes, mitochondrial FO forms membrane-bending dimers. These dimers self-accommodate into long rows at the terminate of the cristae, possibly the outset step of cristae formation.[12] An diminutive model for the dimeric yeast FO region was determined past cryo-EM at an overall resolution of 3.half-dozen Å.[13]
| Subunit | Human Gene |
|---|---|
| a | MT-ATP6 |
| b | ATP5F1 |
| c | ATP5G1, ATP5G2, ATP5G3 |
Binding model [edit]
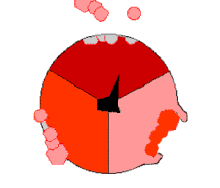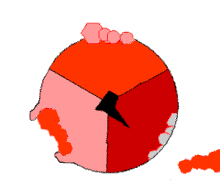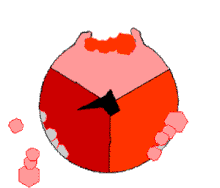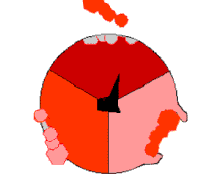
Mechanism of ATP synthase. ADP and Pi (pink) shown being combined into ATP (red), while the rotating γ (gamma) subunit in blackness causes conformational change.

In the 1960s through the 1970s, Paul Boyer, a UCLA Professor, developed the bounden change, or flip-flop, mechanism theory, which postulated that ATP synthesis is dependent on a conformational change in ATP synthase generated by rotation of the gamma subunit. The research grouping of John E. Walker, then at the MRC Laboratory of Molecular Biology in Cambridge, crystallized the Fi catalytic-domain of ATP synthase. The structure, at the time the largest asymmetric protein construction known, indicated that Boyer's rotary-catalysis model was, in essence, correct. For elucidating this, Boyer and Walker shared one-half of the 1997 Nobel Prize in Chemistry.
The crystal construction of the Fone showed alternating alpha and beta subunits (3 of each), arranged like segments of an orangish around a rotating asymmetrical gamma subunit. According to the current model of ATP synthesis (known every bit the alternating catalytic model), the transmembrane potential created past (H+) proton cations supplied by the electron ship chain, drives the (H+) proton cations from the intermembrane infinite through the membrane via the FO region of ATP synthase. A portion of the FO (the ring of c-subunits) rotates as the protons pass through the membrane. The c-ring is tightly attached to the asymmetric cardinal stalk (consisting primarily of the gamma subunit), causing it to rotate within the alphathreebeta3 of Fane causing the 3 catalytic nucleotide binding sites to go through a serial of conformational changes that lead to ATP synthesis. The major F1 subunits are prevented from rotating in sympathy with the key stalk rotor by a peripheral stalk that joins the alphathreebetaiii to the non-rotating portion of FO. The structure of the intact ATP synthase is currently known at low-resolution from electron cryo-microscopy (cryo-EM) studies of the circuitous. The cryo-EM model of ATP synthase suggests that the peripheral stalk is a flexible construction that wraps effectually the circuitous as information technology joins F1 to FO. Under the correct atmospheric condition, the enzyme reaction can too be carried out in reverse, with ATP hydrolysis driving proton pumping across the membrane.
The binding change machinery involves the active site of a β subunit's cycling between iii states.[fourteen] In the "loose" land, ADP and phosphate enter the active site; in the side by side diagram, this is shown in pinkish. The enzyme then undergoes a change in shape and forces these molecules together, with the active site in the resulting "tight" state (shown in red) binding the newly produced ATP molecule with very high affinity. Finally, the active site cycles back to the open state (orange), releasing ATP and binding more than ADP and phosphate, ready for the side by side wheel of ATP production.[fifteen]
Physiological role [edit]
| | This article is missing information almost permeability transition pore (PTP) opening in eukaryotes -- something to do with cell decease and calcium. (January 2022) |
Like other enzymes, the activity of F1FO ATP synthase is reversible. Big-plenty quantities of ATP crusade it to create a transmembrane proton gradient, this is used by fermenting bacteria that do not have an electron transport chain, simply rather hydrolyze ATP to make a proton slope, which they use to drive flagella and the transport of nutrients into the cell.
In respiring bacteria under physiological conditions, ATP synthase, in full general, runs in the contrary direction, creating ATP while using the proton motive strength created by the electron ship concatenation as a source of energy. The overall process of creating free energy in this way is termed oxidative phosphorylation. The same procedure takes place in the mitochondria, where ATP synthase is located in the inner mitochondrial membrane and the F1-office projects into the mitochondrial matrix. The consumption of ATP by ATP-synthase pumps proton cations into the matrix.
Evolution [edit]
The evolution of ATP synthase is idea to have been modular whereby two functionally independent subunits became associated and gained new functionality.[16] [17] This clan appears to have occurred early in evolutionary history, because essentially the aforementioned construction and activity of ATP synthase enzymes are nowadays in all kingdoms of life.[xvi] The F-ATP synthase displays high functional and mechanistic similarity to the V-ATPase.[18] Even so, whereas the F-ATP synthase generates ATP by utilising a proton slope, the V-ATPase generates a proton gradient at the expense of ATP, generating pH values of as low equally 1.[19]
The Fane region also shows pregnant similarity to hexameric Deoxyribonucleic acid helicases (especially the Rho factor), and the entire enzyme region shows some similarity to H +
-powered T3SS or flagellar motor complexes.[eighteen] [20] [21] The α3β3 hexamer of the F1 region shows pregnant structural similarity to hexameric Dna helicases; both form a ring with iii-fold rotational symmetry with a central pore. Both have roles dependent on the relative rotation of a macromolecule within the pore; the Dna helicases employ the helical shape of DNA to drive their motion forth the DNA molecule and to detect supercoiling, whereas the α3βiii hexamer uses the conformational changes through the rotation of the γ subunit to drive an enzymatic reaction.[22]
The H +
motor of the FO particle shows swell functional similarity to the H +
motors that drive flagella.[18] Both characteristic a band of many modest alpha-helical proteins that rotate relative to nearby stationary proteins, using a H +
potential slope every bit an energy source. This link is tenuous, all the same, as the overall structure of flagellar motors is far more complex than that of the FO particle and the band with virtually 30 rotating proteins is far larger than the 10, eleven, or 14 helical proteins in the FO circuitous. More than recent structural data do nevertheless bear witness that the band and the stalk are structurally similar to the F1 particle.[21]
Conformation changes of ATP synthase during synthesis
The modular evolution theory for the origin of ATP synthase suggests that ii subunits with independent function, a DNA helicase with ATPase activity and a H +
motor, were able to bind, and the rotation of the motor collection the ATPase activity of the helicase in reverse.[xvi] [22] This complex then evolved greater efficiency and somewhen developed into today'due south intricate ATP synthases. Alternatively, the Dna helicase/H +
motor circuitous may have had H +
pump action with the ATPase activity of the helicase driving the H +
motor in opposite.[16] This may have evolved to bear out the reverse reaction and act as an ATP synthase.[17] [23] [24]
Inhibitors [edit]
A diverseness of natural and synthetic inhibitors of ATP synthase have been discovered.[25] These take been used to probe the structure and mechanism of ATP synthase. Some may be of therapeutic utilize. There are several classes of ATP synthase inhibitors, including peptide inhibitors, polyphenolic phytochemicals, polyketides, organotin compounds, polyenic α-pyrone derivatives, cationic inhibitors, substrate analogs, amino acid modifiers, and other miscellaneous chemicals.[25] Some of the about commonly used ATP synthase inhibitors are oligomycin and DCCD.
In different organisms [edit]
Leaner [edit]
Eastward. coli ATP synthase is the simplest known form of ATP synthase, with 8 dissimilar subunit types.[11]
Bacterial F-ATPases tin can occasionally operate in reverse, turning them into an ATPase.[26] Some leaner have no F-ATPase, using an A/V-type ATPase bidirectionally.[9]
Yeast [edit]
Yeast ATP synthase is 1 of the best-studied eukaryotic ATP synthases; and five Fane, eight FO subunits, and seven associated proteins accept been identified.[7] Most of these proteins have homologues in other eukaryotes.[27] [28] [29] [30]
Institute [edit]
In plants, ATP synthase is also nowadays in chloroplasts (CF1FO-ATP synthase). The enzyme is integrated into thylakoid membrane; the CF1-office sticks into stroma, where dark reactions of photosynthesis (also called the lite-independent reactions or the Calvin cycle) and ATP synthesis take place. The overall structure and the catalytic machinery of the chloroplast ATP synthase are almost the same as those of the bacterial enzyme. However, in chloroplasts, the proton motive force is generated not by respiratory electron transport chain but by master photosynthetic proteins. The synthase has a 40-aa insert in the gamma-subunit to inhibit wasteful action when dark.[31]
Mammal [edit]
The ATP synthase isolated from bovine (Bos taurus) middle mitochondria is, in terms of biochemistry and construction, the best-characterized ATP synthase. Beef heart is used as a source for the enzyme because of the high concentration of mitochondria in cardiac musculus. Their genes have shut homology to man ATP synthases.[32] [33] [34]
Man genes that encode components of ATP synthases:
- ATP5A1
- ATP5B
- ATP5C1, ATP5D, ATP5E, ATP5F1, ATP5G1, ATP5G2, ATP5G3, ATP5H, ATP5I, ATP5J, ATP5J2, ATP5L, ATP5O
- MT-ATP6, MT-ATP8
Other eukaryotes [edit]
Eukaryotes belonging to some divergent lineages accept very special organizations of the ATP synthase. A euglenozoa ATP synthase forms a dimer with a boomerang-shaped F1 caput like other mitochondrial ATP synthases, but the FO subcomplex has many unique subunits. It uses cardiolipin. The inhibitory IFi also binds differently, in a mode shared with trypanosomatida.[35]
Archaea [edit]
Archaea do non generally accept an F-ATPase. Instead, they synthesize ATP using the A-ATPase/synthase, a rotary machine structurally similar to the 5-ATPase simply mainly functioning as an ATP synthase.[26] Like the bacteria F-ATPase, it is believed to too function as an ATPase.[9]
See also [edit]
- ATP10 poly peptide required for the assembly of the FO sector of the mitochondrial ATPase circuitous.
- Chloroplast
- Electron transfer concatenation
- Flavoprotein
- Mitochondrion
- Oxidative phosphorylation
- P-ATPase
- Proton pump
- Rotating locomotion in living systems
- Transmembrane ATPase
- V-ATPase
References [edit]
- ^ Okuno D, Iino R, Noji H (April 2011). "Rotation and structure of FOF1-ATP synthase". Journal of Biochemistry. 149 (6): 655–64. doi:10.1093/jb/mvr049. PMID 21524994.
- ^ Junge W, Nelson N (June 2015). "ATP synthase". Annual Review of Biochemistry. 84: 631–57. doi:10.1146/annurev-biochem-060614-034124. PMID 25839341.
- ^ Kagawa Y, Racker E (May 1966). "Partial resolution of the enzymes catalyzing oxidative phosphorylation. eight. Properties of a factor conferring oligomycin sensitivity on mitochondrial adenosine triphosphatase". The Journal of Biological Chemistry. 241 (10): 2461–six. doi:10.1016/S0021-9258(18)96640-8. PMID 4223640.
- ^ Mccarty RE (November 1992). "A plant biochemist's view of H +
-ATPases and ATP synthases". The Periodical of Experimental Biology. 172 (Pt i): 431–441. doi:10.1242/jeb.172.1.431. PMID 9874753. - ^ PDB: 5ARA; Zhou A, Rohou A, Schep DG, Bason JV, Montgomery MG, Walker JE, Grigorieff N, Rubinstein JL (Oct 2015). "Structure and conformational states of the bovine mitochondrial ATP synthase past cryo-EM". eLife. 4: e10180. doi:10.7554/eLife.10180. PMC4718723. PMID 26439008.
- ^ Goodsell D (Dec 2005). "ATP Synthase". Molecule of the Month. doi:10.2210/rcsb_pdb/mom_2005_12.
- ^ a b Velours J, Paumard P, Soubannier V, Spannagel C, Vaillier J, Arselin Chiliad, Graves PV (May 2000). "System of the yeast ATP synthase F(0):a study based on cysteine mutants, thiol modification and cantankerous-linking reagents". Biochimica et Biophysica Acta (BBA) - Bioenergetics. 1458 (2–3): 443–56. doi:10.1016/S0005-2728(00)00093-1. PMID 10838057.
- ^ Fernandez Moran H, Oda T, Blair PV, Green DE (July 1964). "A macromolecular repeating unit of mitochondrial structure and office. Correlated electron microscopic and biochemical studies of isolated mitochondria and submitochondrial particles of beefiness heart musculus". The Journal of Cell Biological science. 22 (1): 63–100. doi:10.1083/jcb.22.1.63. PMC2106494. PMID 14195622.
- ^ a b c Stewart AG, Laming EM, Sobti Chiliad, Stock D (April 2014). "Rotary ATPases--dynamic molecular machines". Current Opinion in Structural Biology. 25: 40–8. doi:x.1016/j.sbi.2013.xi.013. PMID 24878343.
- ^ PDB: 1VZS; Carbajo RJ, Silvester JA, Runswick MJ, Walker JE, Neuhaus D (2004). "Solution structure of subunit F(6) from the peripheral stem region of ATP synthase from bovine heart mitochondria". Journal of Molecular Biology. 342 (2): 593–603. doi:10.1016/j.jmb.2004.07.013. PMID 15327958.
- ^ a b Ahmad Z, Okafor F, Laughlin TF (2011). "Role of Charged Residues in the Catalytic Sites of Escherichia coli ATP Synthase". Journal of Amino Acids. 2011: i–12. doi:x.4061/2011/785741. PMC3268026. PMID 22312470.
- ^ Blum TB, Hahn A, Meier T, Davies KM, Kühlbrandt W (March 2019). "Dimers of mitochondrial ATP synthase induce membrane curvature and self-gather into rows". Proceedings of the National University of Sciences of the U.s.a. of America. 116 (10): 4250–4255. doi:10.1073/pnas.1816556116. PMC6410833. PMID 30760595.
- ^ Guo H, Bueler SA, Rubinstein JL (Nov 2017). "Diminutive model for the dimeric FO region of mitochondrial ATP synthase". Science. 358 (6365): 936–40. Bibcode:2017Sci...358..936G. doi:10.1126/science.aao4815. PMC6402782. PMID 29074581.
- ^ Gresser MJ, Myers JA, Boyer PD (Oct 1982). "Catalytic site cooperativity of beef heart mitochondrial F1 adenosine triphosphatase. Correlations of initial velocity, bound intermediate, and oxygen exchange measurements with an alternating three-site model". The Journal of Biological Chemistry. 257 (20): 12030–8. doi:10.1016/S0021-9258(18)33672-X. PMID 6214554.
- ^ Nakamoto RK, Baylis Scanlon JA, Al-Shawi MK (August 2008). "The rotary mechanism of the ATP synthase". Archives of Biochemistry and Biophysics. 476 (i): 43–50. doi:x.1016/j.abb.2008.05.004. PMC2581510. PMID 18515057.
- ^ a b c d Doering C, Ermentrout B, Oster G (Dec 1995). "Rotary DNA motors". Biophysical Periodical. 69 (six): 2256–67. Bibcode:1995BpJ....69.2256D. doi:10.1016/S0006-3495(95)80096-2. PMC1236464. PMID 8599633.
- ^ a b Crofts A. "Lecture 10:ATP synthase". Life Sciences at the Academy of Illinois at Urbana–Champaign.
- ^ a b c "ATP Synthase". InterPro Database.
- ^ Beyenbach KW, Wieczorek H (February 2006). "The V-blazon H+ ATPase: molecular structure and function, physiological roles and regulation". The Journal of Experimental Biology. 209 (Pt 4): 577–89. doi:10.1242/jeb.02014. PMID 16449553.
- ^ Skordalakes East, Berger JM (July 2003). "Construction of the Rho transcription terminator: mechanism of mRNA recognition and helicase loading". Cell. 114 (ane): 135–46. doi:10.1016/S0092-8674(03)00512-9. PMID 12859904. S2CID 5765103.
- ^ a b Imada Thousand, Minamino T, Uchida Y, Kinoshita Chiliad, Namba Thou (March 2016). "Insight into the flagella type III export revealed by the complex construction of the blazon Iii ATPase and its regulator". Proceedings of the National Academy of Sciences of the Us of America. 113 (13): 3633–8. Bibcode:2016PNAS..113.3633I. doi:ten.1073/pnas.1524025113. PMC4822572. PMID 26984495.
- ^ a b Martinez LO, Jacquet Southward, Esteve JP, Rolland C, Cabezón E, Champagne East, et al. (January 2003). "Ectopic beta-chain of ATP synthase is an apolipoprotein A-I receptor in hepatic HDL endocytosis". Nature. 421 (6918): 75–9. Bibcode:2003Natur.421...75M. doi:10.1038/nature01250. PMID 12511957. S2CID 4333137.
- ^ Cross RL, Taiz L (January 1990). "Cistron duplication as a means for altering H+/ATP ratios during the evolution of FOFone ATPases and synthases". FEBS Letters. 259 (2): 227–nine. doi:10.1016/0014-5793(90)80014-a. PMID 2136729. S2CID 32559858.
- ^ Cantankerous RL, Müller 5 (October 2004). "The development of A-, F-, and V-blazon ATP synthases and ATPases: reversals in role and changes in the H+/ATP coupling ratio". FEBS Letters. 576 (1–2): 1–four. doi:10.1016/j.febslet.2004.08.065. PMID 15473999. S2CID 25800744.
- ^ a b Hong S, Pedersen PL (December 2008). "ATP synthase and the actions of inhibitors utilized to study its roles in human health, illness, and other scientific areas". Microbiology and Molecular Biology Reviews. 72 (4): 590–641, Table of Contents. doi:ten.1128/MMBR.00016-08. PMC2593570. PMID 19052322.
- ^ a b Kühlbrandt W, Davies KM (January 2016). "Rotary ATPases: A New Twist to an Ancient Automobile". Trends in Biochemical Sciences. 41 (i): 106–116. doi:ten.1016/j.tibs.2015.x.006. PMID 26671611.
- ^ Devenish RJ, Prescott M, Roucou X, Nagley P (May 2000). "Insights into ATP synthase assembly and function through the molecular genetic manipulation of subunits of the yeast mitochondrial enzyme complex". Biochimica et Biophysica Acta (BBA) - Bioenergetics. 1458 (two–iii): 428–42. doi:10.1016/S0005-2728(00)00092-X. PMID 10838056.
- ^ Kabaleeswaran Five, Puri N, Walker JE, Leslie AG, Mueller DM (November 2006). "Novel features of the rotary catalytic mechanism revealed in the structure of yeast F1 ATPase". The EMBO Journal. 25 (22): 5433–42. doi:x.1038/sj.emboj.7601410. PMC1636620. PMID 17082766.
- ^ Stock D, Leslie AG, Walker JE (November 1999). "Molecular compages of the rotary motor in ATP synthase". Science. 286 (5445): 1700–5. doi:10.1126/science.286.5445.1700. PMID 10576729.
- ^ Liu S, Charlesworth TJ, Bason JV, Montgomery MG, Harbour ME, Fearnley IM, Walker JE (May 2015). "The purification and label of ATP synthase complexes from the mitochondria of 4 fungal species". Biochemical Journal. 468 (1): 167–175. doi:10.1042/BJ20150197. PMC4422255. PMID 25759169.
- ^ Hahn A, Vonck J, Mills DJ, Meier T, Kühlbrandt W (May 2018). "Construction, machinery, and regulation of the chloroplast ATP synthase". Science. 360 (6389): eaat4318. doi:10.1126/science.aat4318. PMC7116070. PMID 29748256.
- ^ Abrahams JP, Leslie AG, Lutter R, Walker JE (August 1994). "Structure at 2.eight A resolution of F1-ATPase from bovine heart mitochondria". Nature. 370 (6491): 621–viii. Bibcode:1994Natur.370..621A. doi:10.1038/370621a0. PMID 8065448. S2CID 4275221.
- ^ Gibbons C, Montgomery MG, Leslie AG, Walker JE (November 2000). "The structure of the central stem in bovine F(1)-ATPase at 2.4 A resolution". Nature Structural Biological science. 7 (11): 1055–61. doi:10.1038/80981. PMID 11062563. S2CID 23229994.
- ^ Menz RI, Walker JE, Leslie AG (August 2001). "Structure of bovine mitochondrial F(ane)-ATPase with nucleotide bound to all three catalytic sites: implications for the mechanism of rotary catalysis". Prison cell. 106 (three): 331–41. doi:x.1016/s0092-8674(01)00452-4. PMID 11509182. S2CID 1266814.
- ^ Mühleip A, McComas SE, Amunts A (November 2019). "Structure of a mitochondrial ATP synthase with bound native cardiolipin". eLife. 8: e51179. doi:ten.7554/eLife.51179. PMC6930080. PMID 31738165.
- "Dissimilar from the remainder". eLife. December 24, 2019.
Further reading [edit]
- Nick Lane: The Vital Question: Energy, Evolution, and the Origins of Complex Life, Ww Norton, 2015-07-20, ISBN 978-0393088816 (Link points to Figure 10 showing model of ATP synthase)
External links [edit]
- Boris A. Feniouk: "ATP synthase — a splendid molecular machine"
- Well illustrated ATP synthase lecture by Antony Crofts of the University of Illinois at Urbana–Champaign.
- Proton and Sodium translocating F-type, V-type and A-blazon ATPases in OPM database
- The Nobel Prize in Chemistry 1997 to Paul D. Boyer and John E. Walker for the enzymatic mechanism of synthesis of ATP; and to Jens C. Skou, for discovery of an ion-transporting enzyme, Na +
, K +
-ATPase. - Harvard Multimedia Production Site — Videos – ATP synthesis blitheness
- David Goodsell: "ATP Synthase- Molecule of the Month"
Source: https://en.wikipedia.org/wiki/ATP_synthase
0 Response to "What protein (enzyme) is used to produce ATP?"
Post a Comment